Advanced Statistical Resampling Techniques for Complex Research Data
As a trusted statistical resampling service provider, Statswork delivers:
- Monte Carlo simulation services
- Bootstrap and jackknife analysis services
- Permutation and randomization tests
- Cross-validation techniques
These statistical resampling techniques are important, especially while handling small samples, non-normal data, high variability, and/or unknown distributions.
Business and Research Applications of Resampling Data Analysis Services
Our resampling statistical consulting supports organizations in:
- Model validation and predictive analytics
- Environmental and spatial data modeling
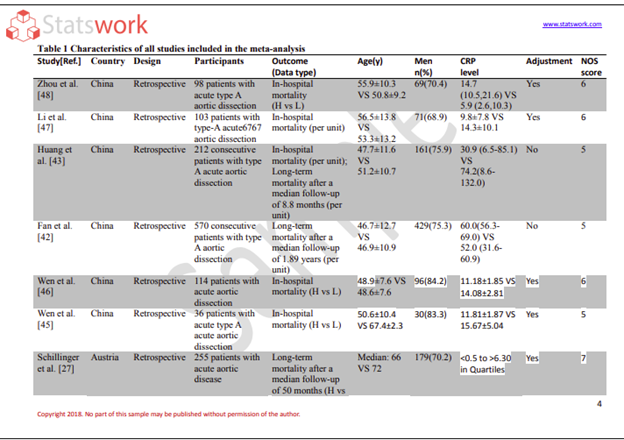
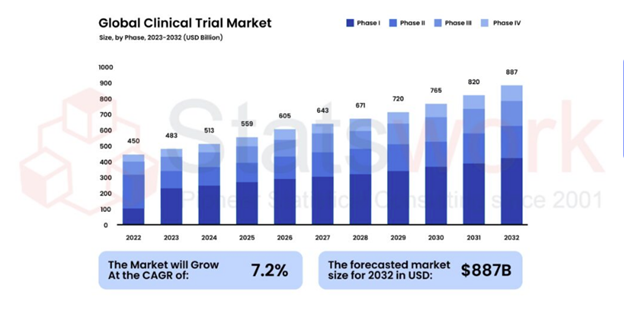
- Healthcare and clinical research analytics
- Market research and behavioral data studies
- Population studies and community structure analysis
- Hypothesis validation in complex datasets
Structured Resampling Workflow Followed by Statswork Experts
Our resampling methodology experts follow a systematic workflow:
- Define the sampling framework and statistical measures
- Use real input datasets for resampling
- Perform repeated sampling with or without replacement
- Compute the required statistical measures
- Run iterations until output distribution stability is achieved
- Generate p-values, confidence intervals, and interpretations
This assures statistically defensible results for organizational research and analytics.
Resampling in R Programming, SPSS, and SAS
We provide:
- Resampling in R programming
- Resampling using SPSS and bootstrap analysis in SPSS
- Resampling using SAS
This enables seamless integration with your existing analytics ecosystem.
Bootstrap and Jackknife Analysis Services for Small Sample Size Studies
Our bootstrap and jackknife analysis services are ideal for:
- Small sample size research
- Bias and variance estimation
- Improving the accuracy of statistical estimates
- Non-parametric statistical validation
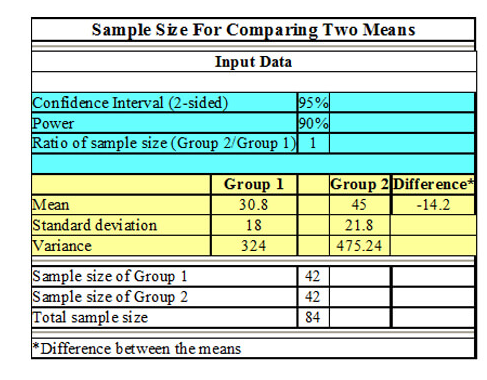
Resampling for Power and Sample Size Calculation
Statswork helps organizations with resampling to determine power and sample size, hence designing studies based on adequate sample size statistics.
Outsourced Resampling Statistical Analysis Services for Organizations
Organizations outsource to Statswork for:
- Professional resampling analysis services
- Resampling statistical consulting
- Resampling techniques for small sample size studies
- Reliable resampling data analysis services