Bioinformatics analysis and identification of potential genes related to pathogenesis
Introduction
Cervical cancer develops over 10 to 25 years, starting with inflammation and progressing via cervical intraepithelial neoplasia (CIN) to invasive malignancy. CIN is a possibly premalignant alteration of the cervix’s squamous cells. Many research has focused on the variety or heterogeneity of distinct solid tumour types in recent years. Cancer cells and tumours have more complicated gene expression network patterns than normal cells and organs. Biological analysis is a scientific methodology that associate analytical tools and biological content in one place so that researchers can obtain a fundamentally deeper and broader understanding of biological relationships and processes
Data Procession
The Gene Expression Omnibus (GEO) database was used to get GSE63514 gene expression profiles. The GSE63514 was a 128-sample expression profiling based on the GPL570 platform (24 normal samples, 76 CIN samples and 28 cervical cancer samples). From a flash-frozen biopsy, all samples were cryosectioned. The array probes were mapped to their particular Gene IDs before bioinformatics investigation using the array annotations. If a probe matches several genes, it will be deleted. If a gene matches several probes, the average value will be calculated. Based on the number of genes filtered out, an appropriate threshold was determined. The study’s workflow is depicted in Figure 1.
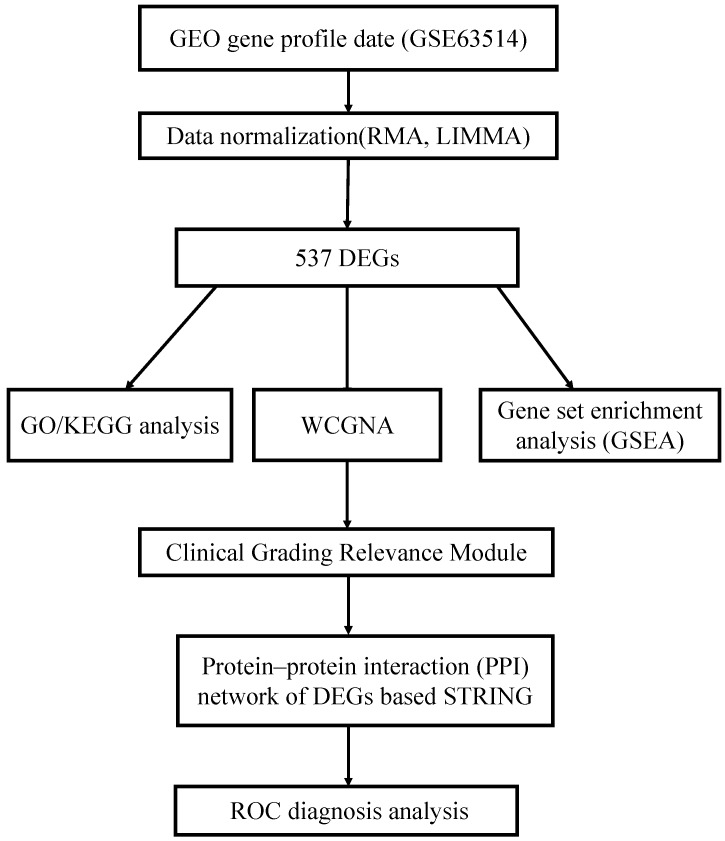
Figure 1. The combined analysis and functional validation flowchart [1].
Gene function and pathways
The Cluster Profiler is an ontology-based R tool that compares gene clusters using biological word categorisation and enrichment analysis better to understand the higher-order functions of biological systems. DAVID -To identify the biological characteristics such as biological process (BP), cellular component (CC), and molecular function (MF) of significant DEGs, a standard functional annotation tool of bioinformatics resources were used. Pathway enrichment analysis from the Kyoto Encyclopedia of Genes and Genomes was utilised to identify the critical pathways. The cut-off threshold for substantial enrichment was established at AdjP-value 0.05.
Gene set enrichment analysis
The enrichment studies were carried out to see if a set of biological processes established in advance were enriched. The enriched pathways were sorted by their normalised enrichment scores (NESs), using a cut-off of FDR 0.05 as the criterion.
Conclusion
DEGs in CIN samples can be utilised to detect the disease’s progression before it progresses to malignancy. Researchers used a combinatorial method that included gene expression profiles, PPI networks, hubs, modules, and motifs to find possible prognostic indicators capable of differentiating progressive cervical illness. Gene expression profiling found 537 DEGs in CIN samples (331 up-regulated genes and 206 down-regulated genes). These genes influenced the cell cycle, DNA replication, Fanconi anaemia route, p53 signalling pathway, homologous recombination, Oocyte meiosis, Mismatch repair, Pyrimidine metabolism, Progesterone-mediated oocyte maturation, and Drug metabolism, among other processes. The four functional gene sets that were enriched were E2F-Targets, G2M-Checkpoint, Mitotic-Spindle, and Spermatogenesis. A total of 31 DEGs were identified as potential hub genes for CIN high-grade risk. Out of 537. 13 genes may interact more closely in CIN categorisation and have a strong diagnostic value. For best clinical research services, you can reach statswok.
Reference
[1] Zhang, X., Bai, J., Yuan, C., Long, L., Zheng, Z., Wang, Q., Chen, F., & Zhou, Y. (2020). Bioinformatics analysis and identification of potential genes related to pathogenesis of cervical intraepithelial neoplasia. Journal of Cancer, 11(8), 2150–2157. https://doi.org/10.7150/jca.38211. [2] Hui Zhang, Jianing Zhong, Youbing Tu, Benquan Liu, Zhibo Chen, Yunchen Luo, Yaping Tang, Fei Xiao, Jincai Zhong, “Integrated Bioinformatics Analysis Identifies Hub Genes Associated with the Pathogenesis and Prognosis of Esophageal Squamous Cell Carcinoma”, BioMed Research International, vol. 2019, Article ID 2615921, 9 pages, 2019. https://doi.org/10.1155/2019/2615921



 Previous Post
Previous Post